 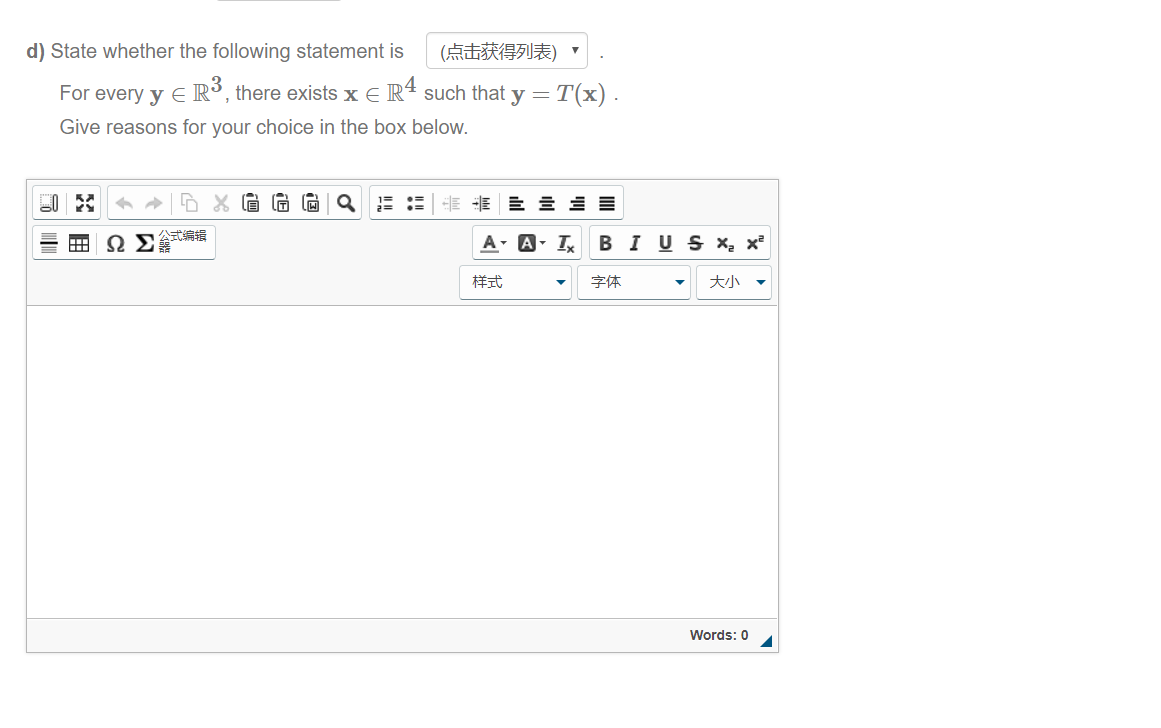
We speculate that the absent MAGEB18 and ERVKs genes may simply be due to the two species’ different evolutionary histories, while the 19 non-coding RNAs may be unrecoverable simply due to their very short lengths. And notably, 26 genes, including MAGEB18, PUF-like, five endogenous retrovirus group K members ( ERVKs), eight snRNA, ten snoRNA, and one tRNA, were completely missed by Christmas Island rat data ( Figure 2 Table S3). These results suggested that most of the long thick black hair, long dark whisker, and round ear phenotypes of the Christmas Island rat could likely be recreated if genome editing of a Norway rat was attempted. Additionally, all eight orthologs of the human round-ear phenotype-involved-genes ( CEP57, ERF, MYH3, NALCN, PSMC3, TNNI2, TNNT3, and TPM2 ) were found to be covered at higher than 97%. 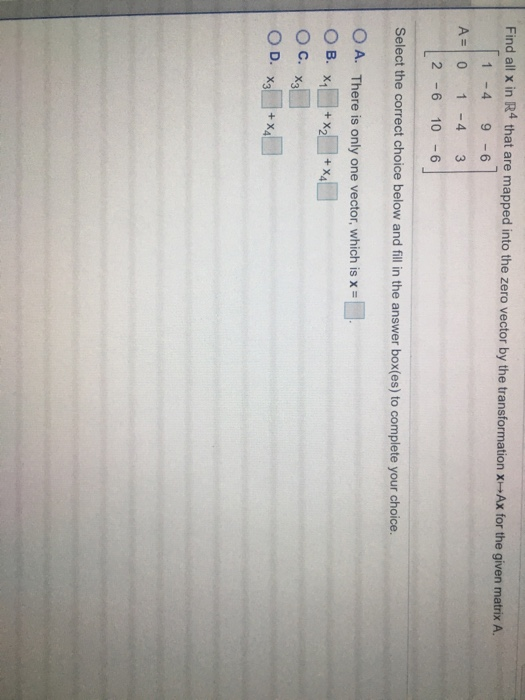
Almost all genes (83/86) encoding keratins or keratin-associated proteins, which are the key structural materials of hair and whiskers, have coverage higher than 90%. We found that 17,121 (50.19%) genes were covered at higher than 0.99 completeness. 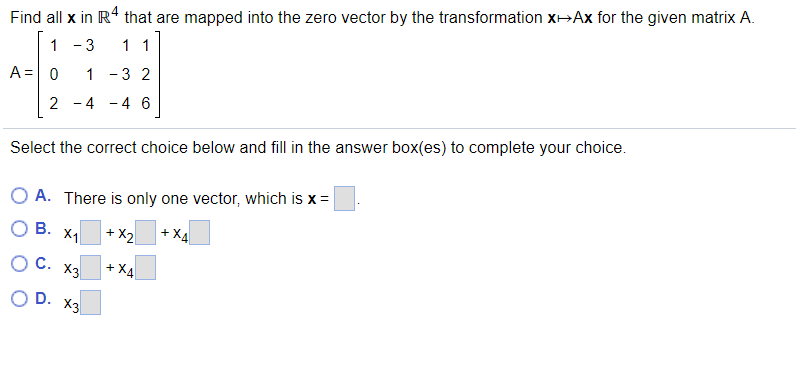
We then calculated the coverage of each of the 34,200 genes annotated in the Norway rat reference genome, including 22,228 protein coding genes and 11,972 non-coding genes ( Figure 2 Table S3). 
one-quarter of it fell within gene regions ( Table S2), thus implying that information is missing that would likely have functional consequences. To answer this question, we explored the genomic distribution of the 128,423,913 bp of the Norway brown rat genome that was not covered by Christmas Island rat sequence data, and found that ca. While this can be partly explained by the observation that 0.81% of the bases in the reference genome are undetermined (Ns), we hypothesized that the remaining 4.04% of the reference genome was unmappable because either (1) the short length of the ancient DNA templates (take the BGISeq data, for example 48.27% of the reads are shorter than 50 bp Figure S1A) reduces their mapping ability (2) the AT richness of some genome regions introduces PCR amplification bias, thus sequencing bias and/or (3) the missing regions are unmappable due to the evolutionary divergence of the two species. Nevertheless, despite this high average depth of coverage, the data only spanned 95.15% of the Norway brown rat reference genome ( Table 1), raising the question as to why. The amount of sequence data generated allowed us to map the Christmas Island rat’s genome sequence to an average depth of 60.81× once the data from the two samples was merged. The sequences displayed characteristic aDNA damage profiles such as misincorporations and fragmentation ( Figure S1 Table S1). Ultimately, our approach demonstrates the importance of applying similar analyses to candidates for de-extinction through genome editing in order to provide critical baseline information about how representative the edited form would be of the extinct species. Furthermore, we find the distribution of regions affected is not random, but for example, if 90% completeness is used as the cutoff, genes related to immune response and olfaction are excessively affected. Our analyses show that even when the extremely high-quality Norway brown rat ( R. norvegicus) is used as a reference, nearly 5% of the genome sequence is unrecoverable, with 1,661 genes recovered at lower than 90% completeness, and 26 completely absent. We then explored how evolutionary divergence from the extant reference genome affected the fraction of the Christmas Island rat genome that could be recovered. It follows that the left null space (the null space of A T) is the orthogonal complement to the column space of A.įor a matrix A, the column space, row space, null space, and left null space are sometimes referred to as the four fundamental subspaces.We first re-sequenced its genome to an average of >60× coverage, then mapped it to the reference genomes of different Rattus species. Thus A T x = 0 if and only if x is orthogonal (perpendicular) to each of the column vectors of A. Let F īecause row vectors of A T are transposes of column vectors v k of A. The column space of a matrix is the image or range of the corresponding matrix transformation. In linear algebra, the column space (also called the range or image) of a matrix A is the span (set of all possible linear combinations) of its column vectors. The column space of this matrix is the vector space spanned by the column vectors. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
